December 31, 2019
Yun Song, Yongjiang Zhang, Jin Xu, Weimin Li & MingFu Li
Scientific Reports volume 9, Article number: 20401 (2019)
Abstract
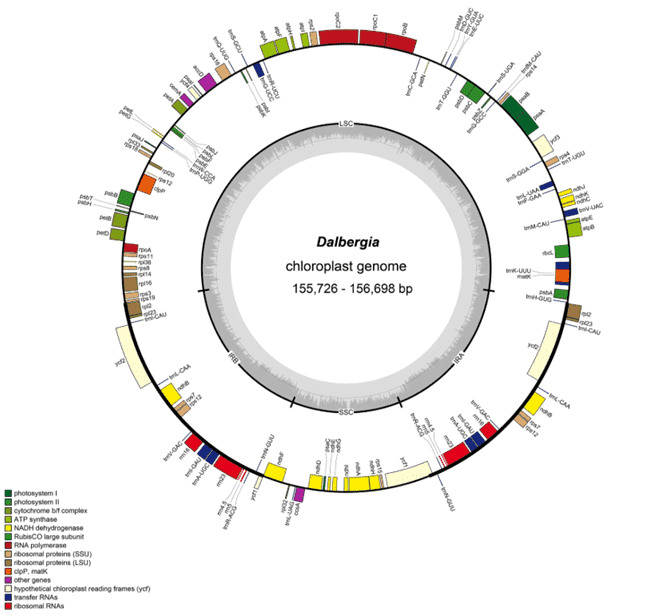
The pantropical plant genus Dalbergia comprises approximately 250 species, most of which have a high economic and ecological value. However, these species are among the most threatened due to illegal logging and the timber trade. To enforce protective legislation and ensure effective conservation of Dalbergia species, the identity of wood being traded must be accurately validated. For the rapid and accurate identification of Dalbergia species and assessment of phylogenetic relationships, it would be highly desirable to develop more effective DNA barcodes for these species. In this study, we sequenced and compared the chloroplast genomes of nine species of Dalbergia. We found that these chloroplast genomes were conserved with respect to genome size, structure, and gene content and showed low sequence divergence. We identified eight mutation hotspots, namely, six intergenic spacer regions (trnL-trnT, atpA-trnG, rps16-accD, petG-psaJ, ndhF-trnL, and ndhG-ndhI) and two coding regions (ycf1a and ycf1b), as candidate DNA barcodes for Dalbergia. Phylogenetic analyses based on whole chloroplast genome data provided the best resolution of Dalbergia, and phylogenetic analysis of the Fabaceae showed that Dalbergia was sister to Arachis. Based on comparison of chloroplast genomes, we identified a set of highly variable markers that can be developed as specific DNA barcodes.
More info at: https://www.nature.com/articles/s41598-019-56727-x
For research purposes only, not intended for clinical diagnosis, treatment, or individual health assessments.


 Sample Submission Guidelines
Sample Submission Guidelines

